P-Curve: A tool for detecting publication bias
How can we tell when publication bias has led to data mining? In this video, we’ll show you what the distribution of p-values should look like when (1) there are no observable effects of a treatment, (2) there are observable effects, and (3) data mining is likely to have occurred. We also discuss a 2014 article authored by Uri Simonsohn, Leif Nelson, and Joseph Simmons (the same authors of “False-Positive Psychology”) in the Journal of Experimental Psychology that demonstrated widespread data mining in that body of literature.
In this article, authors Joseph Simmons, Leif Nelson, and Uri Simonsohn propose a way to distinguish between truly significant findings and false positives resulting from selective reporting and specification searching, or p-hacking.
P values indicate “how likely one is to observe an outcome at least as extreme as the one observed if the studied effect were nonexistent.” As a reminder, most academic journals will only publish studies with p values less than 0.05, the most common threshold for statistical significance.
Some researchers use p-hacking to “find statistically significant support for nonexistent effects,” allowing them to “get most studies to reveal significant relationships between truly unrelated variables.”
The p-curve can be used to detect p-hacking. The authors define this curve as “the distribution of statistically significant p values for a set of independent findings. Its shape is a diagnostic of the evidential value of that set of findings.”
In order for p-curve inferences to be credible, the p values selected must be:
- associated with the hypothesis of interest,
- statistically independent from other selected p values, and
- distributed uniformly under the null.
It is also important to clarify that the p-curve assesses only reported data and not the theories that they are testing. Similarly, it’s important to keep in mind that if a set of values is found to have evidential value, it doesn’t automatically imply internal or external validity.
Using the p-curve to detect p-hacking is fairly straightforward. If the curve is right-skewed as in the chart to the right in the figure below, there are more low (0.01s) than high (0.04s) significant p values, suggesting truly significant p values. When non-existent effects are studied (i.e., a study’s null hypothesis is true), all p values are equally likely to be observed, thus producing a uniform curve or a straight line. In the figure below, each chart incorporates a uniform curve that is dotted and red for comparison. Curves that are left-skewed however, as in the chart to the left in the figure below, indicate more high p values than low ones; p-hacking has likely occurred.
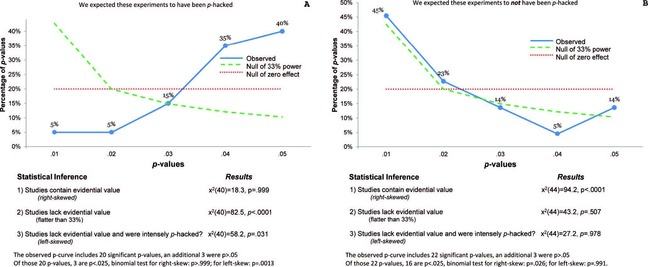 “P-curves for Journal of Personality and Social Psychology” (Click to expand)
“P-curves for Journal of Personality and Social Psychology” (Click to expand)
The above figure displays the results of the authors’ demonstration of the p-curve through the analysis of two sets of findings taken from the Journal of Personality and Social Psychology (JPSP). They hypothesized that one set was p-hacked, while the other was not. In the set in which they suspected p-hacking, they realized that the authors of the publication reported results only with a covariate. While there is nothing wrong with including covariates in study’s design, many researchers will include one only after their initial analysis (without the covariate) was found to be non-significant.
Simmons, Nelson, and Simonsohn provide guidelines to follow when selecting studies to analyze with the p-curve:
- Create a selection rule. Authors should decide in advance which studies to use.
- Disclose the selection rule.
- Maintain robustness to resolutions of ambiguity If it is unclear whether or not a study should be included, authors should report results both with and without that study. This allows readers to see the extent of the influence of these ambiguous cases.
- Replicate single-article p-curves. Because of the risk of cherry-picking single articles, the authors suggest a direct replication of at least one of the studies in the article to improve the credibility of the p-curve.
In addition to these guidelines, Simmons, Nelson, and Simonsohn also provide five steps to ensure that the selected p-values meet the three selection criteria we mentioned earlier:
- Identify researchers’ stated hypothesis and study design.
- Identify the statistical result testing the stated hypothesis.
- Report the statistical results of interest.
- Recompute precise p-values based on reported test statistics. – This has been made easy through an online app, which you can find at http://p-curve.com/.
- Report robustness results of the p-values to your selection rules.
As with anything, the p-curve is not one hundred percent accurate one hundred percent of the time. The validity of the judgments made from a p-curve may depend on “the number of studies being p-curved, their statistical power, and the intensity of p-hacking”. There isn’t much concern over cherry-picking p-curves to ensure the result of a lack of evidential value. However, such a practice can be prevented simply with the disclosure of selections, ambiguity, sample size, and other study details.
Additionally, there are a few limitations with the p-curve. First, it “does not yet technically apply to studies analyzed using discrete test statistics” and is “less likely to conclude data have evidential value when a covariate correlates with the independent variable of interest.” It also has a hard time detecting confounding variables; if there is a real effect, but also mild p-hacking, it usually won’t detect the latter.
Simmons, Nelson, and Simonsohn conclude that, with the examination of a distribution of p-values, one will be able to identify whether selective reporting was used or not. What do you think about the p-curve? Would you use this tool?
We’ll use http://p-curve.com/ in an Exercise later in this Activity.
You can read the entire paper here.
Reference
Simonsohn, Uri, Leif D. Nelson, and Joseph P. Simmons. 2014. “P-Curve: A Key to the File-Drawer.” Journal of Experimental Psychology: General 143 (2): 534–47. doi:10.1037/a0033242.
© Center for Effective Global Action.